Currently not accepting new graduate students
Honors and Awards:
Doctorate of Science (D.Sc.), Magdalen College, University of Oxford, UK in recognition of seminal biochemical research, and dedication and devotion to teaching, science-communication, and mentorship (2020); Wharton Teaching Excellence Award, Wharton School, University of Pennsylvania, Philadelphia, PA, USA (2019); Award for Excellence in Biology Teaching, University of Pennsylvania, Philadelphia, PA, USA (2019); Christian R. and Mary F. Lindback Foundation Award for Distinguished Teaching (2014); Elected as Fellow of American Association for the Advancement of Science for outstanding fundamental research discoveries on the membrane transport and detoxification of xenobiotics, and for distinguished accomplishments and creativity in science education (2013); Award for Excellence in Biology Teaching, University of Pennsylvania, Philadelphia, PA, USA (2013); Cozzarelli Prize, National Academies of Science (2010) for publication of paper of outstanding scientific excellence and originality; Rebecka and Arie Belldegrun Distinguished Directorship, Roy and Diana Vagelos Life Sciences & Management Program, University of Pennsylvania, USA (2009); Ira H. Abrams Memorial Award for Distinguished Teaching, School of Arts & Science, University of Pennsylvania, Philadelphia, PA, USA (2009); National Academies Fellowship in Life Sciences Education, National Academies, Washington, DC, USA (2009); Award for Excellence in Biology Teaching, University of Pennsylvania, Philadelphia, PA, USA (2005); Award for Excellence in Biology Teaching, University of Pennsylvania, Philadelphia, PA, USA (1996); President's Medal, Society for Experimental Biology, UK for pioneering investigations of primary proton pumps (1990)
D.Sc., Magdalen College, University of Oxford
D.Phil., Plant Biochemistry, University of Oxford
B.Sc., First Class Honors, Biological Sciences, University of Sussex
Biochemistry; Plant Biology; Science Communication; Life Sciences & Management
Our interests have their origins in primary research of the molecular biology, cellular biochemistry and proteomics of vacuolar function with special emphasis on membrane transport proteins and the enzymatic machinery responsible for the detoxification of xenobiotics, especially heavy metals. The long-term objectives of these investigations were to identify the proteins concerned and elucidate their mechanisms of action and regulatory characteristics. The approach taken was that of the 'basic biologist' – the search for general principles, regardless of the organism in which they are to be elucidated, not just principles applicable to plants. Many of the investigations conducted therefore entailed parallel molecular and biochemical manipulations of several model systems including the plant Arabidopsis thaliana, the yeast Saccharomyces cerevisiae and the nematode worm Caenorhabditis elegans. It is by this approach that we were able to make fundamental contributions toward understanding a remarkably broad range of transport and related phenomena of general significance. Working from this background in primary research, in combination with an increased involvement with the communication of science to the educated lay public, our current research efforts are directed toward explaining some of the largely untold stories behind discoveries in the life sciences that have had or may prove to have an unprecedented impact on our lives or the lives of others.
Current Research
Our secondary research efforts are concerned with the interface between the life sciences and their implementation; the difficult transition from discovery in the laboratory to success in the market and/or toward the expansion of humanitarian efforts. To date, we have published four feature articles in this area. The first of these articles, Statins: from fungus to pharma, was largely derived from our teaching activities for the Roy and Diana Vagelos Program in Life Sciences & Management. The model we had in mind was a historical scientific narrative on the discovery and implementation of the statins as drugs for the prevention and treatment of cardiovascular disease (CVD). The impetus for preparing the statins article was a sense that material of this nature would be of immense interest to educated members of the general public because it has something for everyone. It is an example of how our understanding of CVD has undergone radical revision, and in so doing given us a better understanding of how statins do what they do (something that would not have happened if not for the introduction of these drugs); how a serendipitous discovery with striking parallels at all levels to the discovery of the penicillins is quite possibly one of the most significant biomedical accomplishments of the twentieth century; how the convergence and application of basic but disparate cellular biochemical concepts and methodologies spawned one of the best selling drugs; how the original discovery of a drug, or class of drugs, made by one company, required the engagement of another company, either as competitor or collaborator, to bring the drug in question to market; how the juxtaposition of economic with biomedical imperatives can be the deciding factor in determining whether to aggressively push for the implementation of a fundamental discovery; how "plan B" compounds can end up being billion dollar pills (Lipitor in this case). The need for the second feature article, Ivermectin and river blindness, came from the realization that despite the immensity of the river blindness problem very few of us in this part of the world know of the existence of this disease, and even fewer know of the connection between it and something that most of us know something about, the "deworming tablets" given to pets and livestock to protect them from heartworm and similar parasitic infections. Yet, the fact of the matter is that if your dog has been given preventative medication for heartworm it was almost certainly given the very same drug, ivermectin, that has and continues to be used to treat literally tens of millions of people in the developing world; people who would otherwise have to live lives of interminable suffering and anguish. The third feature article, Can skinny fat beat obesity?, is an up to date account of the roles played by brown and beige fat (‘skinny fat’) in keeping white fat, a surplus of which is associated with cardiovascular disease, type 2 diabetes and the metabolic syndrome, at bay. Whether the readers are themselves overweight or know others who are, this article is of general interest because it encompasses several unprecedented discoveries made only in the last few years (some only in the last few months) which when explained provide readers with a platform for better understanding the role played by classical brown fat in newborns and hibernating mammals, how beige fat was discovered in animals and human adults, what it is and does, the biochemical basis of thermogenesis, and recent advances in the identification of a new class of therapeutic agents that might eventually be used to combat obesity. The fourth feature article, Metformin: out of backwaters and into the mainstream, is especially intriguing because it deals with a drug that has come to assume prominence despite a checkered history. Its story has many ramifications, and is filled with delays, uncertainties and dead ends, as well as fortuitous accidents. Metformin is a drug, born of folklore, with reasonably well-established clinical benefits whose precise mechanism of action has resisted definition. However, that is not to say that it is prescribed only to a select few, nor that it is a structurally sophisticated compound, because both are very far from the case. Metformin is the current standard of care for the treatment of one of the most common chronic conditions in the modern world – type 2 diabetes – and its structure is remarkably simple by comparison with that of many other drugs. It is a modest methylated biguanide (dimethybiguanide, alias N,N-dimethylimidodicarbonimidic diamide) with a molecular weight (165.6 daltons) and structural complexity akin to that of glucose (molecular weight 180.3 daltons). Of the many baffling aspects of the metformin story is the haphazard way in which it first got noticed and the fact that it came to assume the standing it now has at very different times – times separated by decades – in different parts of the developed world. Unforeseen too is the extent to which its applications have since expanded from the treatment of type 2 diabetes to the treatment of prediabetes and polycystic ovary syndrome (PCOS), and even cancer. A number of factors influenced metformin’s uncertain rise to fame, most notably: the landmark discovery of insulin in the early 1920s; the disruptive effects of the First and Second World Wars on research efforts; and the aftershock of fear among American patients when another biguanide, phenformin, was proven to be harmful, after it had been aggressively marketed and prescribed throughout the 1960s and 70s. In a similar vein, one of the most recent feature articles in this series, How glyphosate cropped up, is concerned with the most important herbicide of all time based on the billions of kilograms that have been applied to croplands worldwide. Yet what could be more improbable a scenario in the light of the success of this herbicide, better known as Roundup, than its accidental discovery, as a modified amino acid no bigger than glucose, with a structure so refined that barely any of its substituents are superfluous, that targets with catastrophic consequences an enzyme in a crucial pathway, the shikimate pathway, that we lack but which is indispensable for plant function; chance discoveries that were then to be followed by the equally accidental discovery of a bacterium harboring a version of the target enzyme which by virtue of a single methyl group is resistant to the herbicide in question? Because, in a nutshell, that’s the Roundup/Roundup Ready story. At just about every step along the way the scientific significance of what had been discovered and its mechanistic basis was not understood until long after the fact.
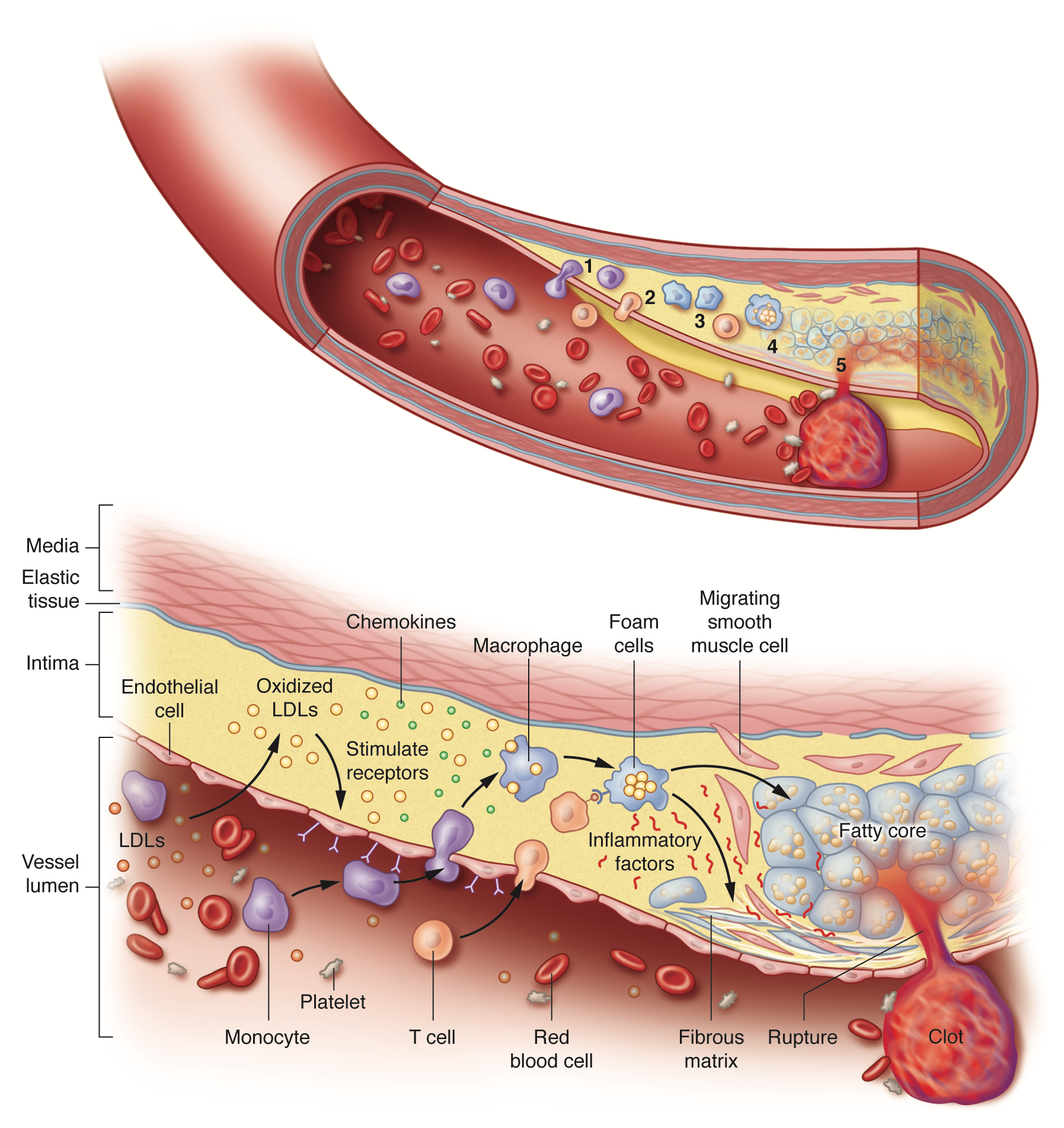
Initiation and progression of atherosclerosis. Atherosclerosis arises from a cascade of biochemical and cellular processes in which low-density lipoproteins (LDLs) trigger inflammation. When their levels in the blood are excessive, LDLs infiltrate the arterial wall where they accumulate and undergo chemical modifications, especially oxidation. The modified LDLs then stimulate the expression of receptors on the innermost, or endothelial, cells lining the artery. In the blood, monocytes – immune cells that participate in general inflammatory responses – dock onto these receptors and enter the arterial wall. Inside the arterial wall, the monocytes mature into macrophages (a class of immune white cells) that engulf the modified LDLs to form fat-filled macrophages called foam cells. The foam cells secrete inflammatory substances that promote the production of a tough fibrous matrix that caps the fatty core to generate a plaque. Acute problems arise when the inflammatory substances secreted by the foam cells weaken the cap. If the cap is breached, blood enters, makes contact with foam-cell proteins that promote clotting and a blood clot develops in the artery. The clot that forms can plug the artery at the site of the plaque or travel downstream and obstruct blood flow at another location. Artist: Sara Jarret (© 2017 Sara Jarret CMI).
An extension of these research activities is the book Managing Discovery in the Life Sciences: Harnessing Creativity to Drive Biomedical Innovation (2018, Cambridge University Press). In this book co-authored with Mark V. Pauly and Lawton R. Burns, case studies of biomedical innovations are presented whereby the reader comes to better understand how the science actually played out through the interplay of personalities and cultures within and between academic and corporate entities and the significance of serendipity not as a mysterious phenomenon but one that is intrinsic to the successes and failures of the experimental approach. The fundamental economic underpinnings of investor-driven discovery management are considered, not as an obstacle or deficiency as its critics would contend or as something beyond reproach as some of its proponents might claim, but as the only means by which scientists and managers can navigate the unknowable to discover new products and decide how to sell them so as to maximize the likelihood of establishing a sustainable pipeline for still more marketable biomedical innovations.
Previous Research
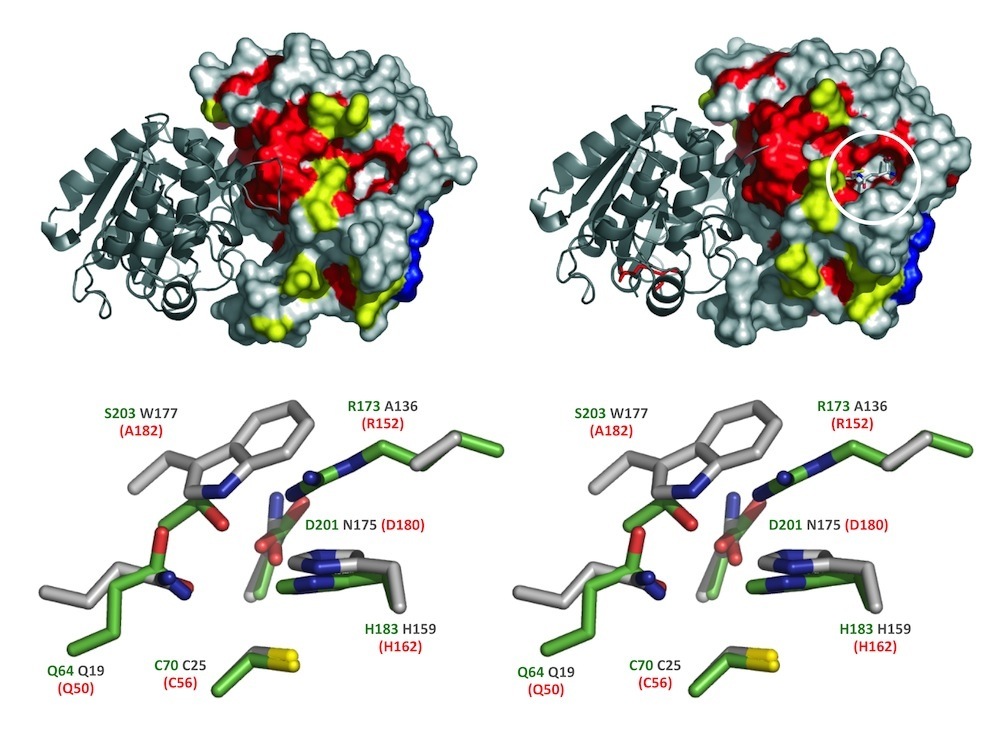
a. phytochelatin-dependent heavy metal detoxification - It has been known for some time that plants and some fungi synthesize peptides termed phytochelatins (PCs) from glutathione (GSH) when exposed to heavy metals, and that PC thiols coordinate and chelate heavy metals to promote their removal from the cytosol by vacuolar sequestration. However, the molecular identity of the enzyme(s) responsible eluded definition until the first cloning of the enzyme PC synthase (AtPCS1) from Arabidopsis by ourselves and two other groups. The isolation of AtPCS1 and its demonstrated sufficiency for PC synthesis from GSH both in vitro and in vivo has enabled detailed mechanistic analyses of this enzyme, and the provision of probes and methodologies for the identification and characterization of its orthologs in animals, as exemplified by our studies of C. elegans. Although studies of PC synthases have largely been concerned with the enzymes from eukaryotes, recent database searches have disclosed PC synthase-like sequences in the genomes of several prokaryotes. This is of particular interest in that all of the prokaryotic PC synthase homologs identified are half the length of their cognates from eukaryotes (220–237 residues compared with 421–506 residues) because they lack the more sequence-variable C-terminal domain. The one prokaryotic PC synthase homolog to have been assayed for activity, the alr0975 protein from the cyanobacterium Nostoc sp. PCC 7120 (NsPCS), catalyzes the deglycylation of GSH to γ-Glu-Cys at a high rate and the synthesis of PC2 at a relatively low rate. A recent crystal structure of NsPCS in its native and γ-Glu-Cys-acylated state (Vivares et al 2006) establishes, as had been inferred from our detailed kinetic, protein chemical, and site mutagenic analyses of the prototypical eukaryotic PC synthase, AtPCS1, that these enzymes belong to the papain superfamily and deploy a cysteine protease-like catalytic mechanism. When account is taken of the fact that in many plant species, PCs are the major ligands responsible for heavy metal accumulation and detoxification, AtPCS1 and genes like it have come to assume prominence in the development of phytoremediation technologies through genetic manipulation of the capacity of plants for heavy metal hyperaccumulation (Rea et al – US Patent No. 6,489,537; Rea et al – US Patent 10,370,675). By the same token, the operation of equivalent metal detoxification pathways in nematodes and their cousins, some of which are pathogenic, but not in vertebrates has spawned research into the development of new strategies centered on this pathway for combating the many human and veterinary diseases caused by these organisms.
Phytochelatin (PC) synthase (PCS) in stereo. PC synthase catalyzes polymerization of the ubiquitous tripeptide, glutathione (γ-Glu-Cys-Gly), to generate short-chain peptides that bind heavy metals with high affinity and contribute to their cellular detoxification. Shown are the crystal structure of a cyanobacterial PCS-like polypeptide (NsPCS) and the identity of the amino acid residues that directly participate in catalysis in NsPCS (green text), its equivalent from the model plant Arabidopsis (red text), and their long-lost distant cousin, the cysteine protease papain (gray text). The active site residues shown are those identified by Vatamaniuk et al (2004), Rea (2006) and Romanyuk et al (2006); the structures are those provided courtesy of Pascal Arnoux and David Pignol and adapted from Vivares et al (2005).
b. ABC transporters - The ATP-binding cassette (ABC) protein superfamily is one the largest protein families known, and most, but not all, are membrane proteins (‘ABC transporters’) active in the transport of a broad range of substances across membranes. Many of these proteins have been implicated in human diseases such as cystic fibrosis and Tangier disease, and resistance to therapies for cancer, malaria, and AIDS. We have played a central role in pioneering investigations of plant ABC transporters and in the discovery of new ABC transporter-mediated heavy metal and organic xenobiotic detoxification pathways through the identification and molecular characterization of glutathione S-conjugate pumps (GS-X pumps) in yeast and plants. GS-X pumps, which belong to the multidrug resistance-associated protein (MRP/ABCC) subfamily of ABC transporters, are involved in the vacuolar sequestration and/or plasma membrane extrusion and detoxification of both endogenous and exogenous toxins. Examples of substances transported by these membrane proteins are herbicides and heavy metals, as exemplified by cadmium and arsenic after their complexation with glutathione (GSH). As such, these transporters and their genes are of potential value for engineering plants with an increased capacity for the removal of toxic materials from contaminated soils and ground waters (Rea et al – US Patent No. 6,166,299). Our subsequent research in this area has been concerned with the molecular and biochemical analysis of plant ABC transporters implicated in the transport of folate (vitamin B9) and its derivatives, and other members of the superfamily implicated in the transport of iron-sulfur clusters across membranes for assembly of the prosthetic groups of oxidoreductases. The overall impact of this work is illustrated by the fact that Arabidopsis allocates a large fraction of its open reading frames (a minimum of 0.5%) to members of the ABC superfamily. To our initial surprise when we compiled the first comprehensive inventory of Arabidopsis ABC proteins, the genome of this organism contains more than 130 ORFs for these proteins, of which more than 100 are transmembrane proteins. This gene count far outstrips those of the genomes of humans and other animals by a wide margin.
c. pyrophosphate-energized proton pumps - Our studies have been instrumental in the discovery and elucidation of the basic organization and core catalytic capabilities of proton-translocating inorganic pyrophosphatases (V-PPases), a novel class of proton pump. Membrane-associated proton-translocating PPases are primary proton pumps that use inorganic pyrophosphate (PPi), the limiting case of a high energy phosphate, instead of ATP as an energy source for the establishment of transmembrane electrochemical potentials . Although the initial objective of these studies was to understand the mechanism of this pump in plants, recent investigations in ours and other laboratories have demonstrated these pumps in organisms as disparate as thermophilic Archaea and parasitic protists. Among the many evolutionary, practical and bioenergetic implications of these findings is the possibility that this research will spawn new approaches to the treatment of several prolific and debilitating parasite-mediated infections. It was originally thought that all V-PPases are of only one type, the type to which the first V-PPase to be cloned belongs. It is now clear that is not the case: we have shown that V-PPases fall into two clearly delineated clades known as the type I and type II enzymes. Type I and type II V-PPases are readily distinguished from each other on the basis of their sequences and the near obligate requirement of the former but not the latter for potassium for activity. In Arabidopsis three genes, AVP1, AVP2, and AVP3 encode V-PPases. AVP1 is the prototypical potassium-dependent type I V-PPase which is expressed ubiquitously and at particularly high levels in developing tissues. AVP2 and AVP3 are the type II V-PPases which share only 36% sequence identity with AVP1 but greater than 85% sequence identity with each other.
d. vacuolar proteomics - The vacuole of S. cerevisiae, which can occupy as much as 25% of total intracellular volume, participates in numerous cellular processes ranging from macromolecule degradation and salvage, pH and general ion homeostasis, osmoregulation and volume regulation, the storage of amino acids, carboxylic acids, carbohydrates and some vitamins, to the sequestration of endogenous and exogenous toxins. What is perhaps surprising given this multifunctionality is how little was known of the range of proteins found in this compartment and the types of modifications to which they are subject. For instance, while protein turnover is one of the most thoroughly investigated functions of the yeast vacuole, and many of the proteins that participate have been known for some time, this and related processes have yet to be explored at the systems level through the application of global proteomics approaches. Using high-purity ‘proteomics-grade’ intact yeast vacuoles, we have reproducibly resolved 360 luminal polypeptide species by two-dimensional gel electrophoresis . Of these, 117 have been identified by MALDI-TOF MS and/or LCQ-MS through the deployment of ProteinLynx-, MASCOT- and/or SEQUEST-based protein sequence database searches in combination with Mr and pI considerations. The polypeptides identified, many of which correspond to alternate isoelectric and size states of the same parent translation product, can be assigned to 66 unique reading frames. In strict agreement with a predominantly lysosomal function for the yeast vacuole, most of the proteins identified are either canonical vacuolar proteases or proteins involved in intermediary metabolism, protein synthesis, folding or targeting, or the alleviation of oxidative stress that have entered this compartment for salvage purposes.
BIOL 2810 (formerly 204) - Biochemistry
LSMP 1210 (formerly 121) - Proseminar in Management & The Life Sciences
For brief account of teaching philosophy go to http://www.upenn.edu/almanac/volumes/v55/n26/tatl.html
Rea, P.A. (2023) Wie Glyphosat die Welt eroberte. Spektrum der Wissenschaft, www.spektrum.de/artikel/2169858,56-63.
Rea, P.A. (2022) How glyphosate cropped up. Am. Sci., 110: 170-177.
Rea, P.A. (2020) Phytochelatin synthase. In: Encyclopedia of Life Sciences (eLS). John Wiley & Sons, Ltd: Chichester, DOI: 10.1002/9780470015902.a0028220, 1-15.
Rea P.A., Pauly, M.V., Burns, L.R. (2018) Managing Discovery in the Life Sciences: Harnessing Creativity to Drive Biomedical Innovation. Cambridge: Cambridge University Press, ISBN 9781316459263, 542 pages.
Rea, P.A. (2018) Plant Vacuoles. In: Encyclopedia of Life Sciences (eLS). John Wiley & Sons, Ltd: Chichester, DOE: 10.10-02/9780470015902.a0001675.pub3, 1-14
Rea, P.A., Tien, A.Y. (2017) Metformin: out of backwaters and into the mainstream. Am. Sci., 105: 102-111.
Rea, P.A., Yin, P., Zahalka, R. (2015) Mit beigem Fett gegen Übergewicht? Spektrum der Wissenschaft, www.spektrum.de/artikel/1346952, 26-32.
Cahoon, R.E., Lutke, W.K., Cameron, J.C., Chen, S., Lee, S.G., Rivard, R.S., Rea, P.A., Jez, J.M. (2015) Adaptive engineering of phytochelatin-based heavy metal tolerance. J. Biol. Chem., 290: 17321-17330.
Rea, P.A., Yin, P., Zahalka, R. (2014) Chew the fat. Am. Sci., 102: 325.
Rea, P.A., Yin, P., Zahalka, R. (2014) Can skinny fat beat obesity? Am. Sci., 102: 272-279.
Rea, P.A. (2012) Phytochelatin synthase: of a protease a peptide polymerase made. Physiol. Plant., 145: 154-164.
Song, W.Y, Park, J., Mendoza-Cózatl, D.G., Suter-Grotemeyer, M., Shim, D., Hörtensteiner, S., Geisler, M., Weder, B., Rea, P.A., Rentsch, D., Schroeder, J.I., Lee, Y., Martinoia, E. (2010) Arsenic tolerance in Arabidopsis is mediated by two ABCC-type phytochelatin transporters. Proc. Natl. Acad. Sci. USA, 107: 21187-21192.
Rea, P.A., Zhang, V., Baras, Y.S. (2010) Ivermectin and river blindness. Am. Sci., 98: 294-303.
Rea, P.A. (2009) TALK ABOUT TEACHING AND LEARNING: The kick is in finding out stuff about stuff and sharing it with others. Almanac, 55: 8.
Raichaudhuri, A., Peng, M., Naponelli, V., Chen, S., Sánchez-Fernández, R., Gu, H., Gregory III, J.F., Hanson, A.D., Rea, P.A. (2009) Plant vacuolar ABC transporters that translocate folates and antifolates in vitro and contribute to antifolate tolerance in vivo. J. Biol. Chem., 284: 8449-8460.
Sooksa-nguan, T., Yakubov, B., Kozlovskyy, V.I., Barkume, C.M., Howe, K.J., Thannhauser, T.W., Rutzke, M.A., Hart, J.J., Kochian, K.V., Rea, P.A., Vatamaniuk, O.K. (2009) Drosophila ABC transporter, DmHMT-1, confers tolerance to cadmium. DmHMT-1 and its yeast homolog, SpHMT-1, are not essential for vacuolar phytochelatin sequestration. J. Biol. Chem., 284: 354-62.
Rea, P.A. (2008) Statins: from fungus to pharma. Am. Sci., 96: 408-415.
Verrier, P.J., Bird, D., Burla, B., Dassa, E., Forestier, C., Geisler, M., Klein, M., Kolukisaoglu, U., Lee, Y., Martinoia, E., Murphy, A., Rea, P.A., Samuels, L., Schulz, B., Spalding, E., Yazaki, K., Theodoulou, F.L. (2008) Plant ABC proteins – a unified nomenclature and updated inventory. Trends Plant Sci., 13: 151-159.
Sarry, J.-E., Chen, S., Collum, R.P., Liang, S., Peng, M., Lang, A., Naumann, B., Dzierszinski, F., Yuan, C.-X., Hippler, M., Rea, P.A. (2007) Analysis of the vacuolar luminal proteome of Saccharomyces cerevisiae. FEBS J., 274: 4287-4305.
Chen, S., Sánchez-Fernández, R., Lyver, E.R., Dancis, A., Rea. P.A. (2007) Functional characterization of AtATM1, AtATM2 and AtATM3, a subfamily of Arabidopsis half-molecule ABC transporters implicated in iron homeostasis. J. Biol. Chem., 282: 21561-21571.
Rea, P.A. (2007) Plant ATP-binding cassette transporters. Annu. Rev. Plant Biol., 58: 347-375.
Romanyuk, N.D., Rigden, D.J., Vatamaniuk, O.K., Lang, A., Cahoon, R.E., Jez, J.M., Rea, P.A. (2006) Mutagenic definition of papain-like catalytic triad, sufficiency of N-terminal domain for single-site core catalytic enzyme acylation and C-terminal domain for augmentative metal activation of an eukaryotic phytochelatin synthase. Plant Physiol., 141:858-869.
Rea, P.A. (2006) Phytochelatin synthase, papain's cousin, in stereo. Proc. Natl. Acad. Sci. USA, 103: 507-508.
Rea, P.A. (2005) A farewell to bacterial ARMs? Nature Biotechnol., 23: 1085-1087.
Orsomando, G., Diaz de la Garza, R., Green, B.J., Peng, M., Rea, P.A., Ryan, T.J., Gregory, J.F., Hanson, A.D. (2005) Plant γ-glutamyl hydrolases and folate polyglutamates. Characterization, compartmentation and co-occurrence in vacuoles. J. Biol. Chem., 280: 28877-28884.
Vatamaniuk, O.K., Bucher, E.A., Sundaram, M.V., Rea, P.A. (2005) CeHMT-1, a putative phytochelatin transporter, is required for cadmium tolerance in Caenorhabditis elegans. J. Biol. Chem., 280: 23684-23690.
Fall (2004) Photographs by Christopher Griffith, Verse by Walt Whitman, Text by Philip A. Rea. Hardcover, 11.25 x 14.25 inches, 80 pages, 48 four-color photographs. Powerhouse Publishers, New York. ISBN 1-57687-226-2.
Rea, P.A., Vatamaniuk, O.K., Rigden, D.J. (2004) Weeds, worms and more: papain's long-lost cousin, phytochelatin synthase. Plant Physiol., 136: 2463-2474.
Vatamaniuk, O.K., Mari, S., Lang, A., Chalasani, S., Demkiv, L.O., Rea, P.A. (2004) Phytochelatin synthase, a dipeptidyl transferase that undergoes multisite acylation with γ-glutamylcysteine during catalysis. Stoichiometric and site-directed mutagenic analysis of AtPCS1-catalyzed phytochelatin synthesis. J. Biol. Chem., 279: 22449-22460.
Rea, P.A. (2003) Ion Genomics. Nature Biotechnol., 21: 1149-1151.
Maathuis, F.J.M., Filatov, V., Krijger, G.C., Herzyk, P., Axelsen, K.B., Chen, S., Green, B.J., Madagan, K.L., Sánchez-Fernández, R., Forde, B., Palmgren, M.G., Rea, P.A., Williams, L.E., Sanders, D., Amtmann, A. (2003) Transcriptome analysis of Arabidopsis thaliana cation transport. Plant J., 65, 675-692.
Rea, P.A., Sánchez-Fernández, R., Chen, S., Peng, M., Klein, M., Geisler, M., Martinoia, M. (2003) The plant ABC transporter superfamily: the functions of a few and the identities of many. In: ABC Transporters from Bacteria to Humans, (Cole, S.P., Kuchler, K., Higgins, C, Holland, B., eds), Academic Press, UK, pp. 335-356.
Drozdowicz, Y.M., Shaw, M., Nishi, M., Striepen, B., Liwinski, H.A., Roos, D.S., and Rea, P.A. (2003) Isolation and characterization of TgVP1, a type I vacuolar proton translocating pyrophosphatase from Toxoplasma gondii: the dynamics of its subcellular localization and the cellular effects of a diphosphonate. J. Biol. Chem., 278: 1075-1085.
Bartholomew, D.M., Van Dyk, D.E., Lau, S.-M., O'Keefe, D.P., Rea, P.A., and Viitanen, P.V. (2002) Alternate energy-dependent pathways for the vacuolar uptake of glucose and glutathione conjugates. High sensitivity, high fidelity transport measurements by LC-MS. Plant Physiol., 103: 1562-1572
Vatamaniuk, O.K., Bucher, E.A., and Rea, P.A. (2002) Worms take the 'phyto' out of 'phytochelatins'. Trends Biotechnol., 20: 61-64
Sánchez-Fernández, R., Davies, T.G.E., Coleman, J.O.D., and Rea, P.A. (2001) The Arabidopsis thaliana ABC protein superfamily: a complete inventory. J. Biol. Chem., 276: 30231-30244
Vatamaniuk, O.K., Bucher, E.A., Ward, J.T., and Rea, P.A. (2001) A new pathway for heavy metal detoxification in animals: phytochelatin synthase is required for cadmium tolerance in Caenorhabditis elegans. J. Biol. Chem., 276: 20817-20820
Drozdowicz, Y.M., and Rea, P.A. (2001) Vacuolar proton-pyrophosphatases: from the evolutionary backwaters into the mainstream. Trends Plant Sci., 6: 206-211
Liu, G., Sánchez-Fernández, R., and Rea, P.A. (2001) Enhanced multispecificity of vacuolar membrane-localized ABC transporter AtMRP2. J. Biol. Chem., 276: 8648-8656
Vatamaniuk, O.K., Mari, S., Lu, Y.-P., and Rea, P.A. (2000) Mechanism of heavy metal activation of phytochelatin (PC) synthase: blocked thiols are sufficient for PC synthase-catalyzed transpeptidation of glutathione and related thiol peptides. J. Biol. Chem., 275: 31451-31459
Drozdowicz, Y.M., Kissinger, J.C., and Rea, P.A. (2000) AVP2, a sequence-divergent, monovalent cation-insensitive proton-translocating inorganic pyrophosphatase from Arabidopsis thaliana. Plant Physiol., 123: 353-362
Drozdowicz, Y.M., Lu, Y.-P., Patel, V., Fitz-Gibbon, S., Miller, J., and Rea, P.A. (1999) PVP, a thermostable vacuolar-type pyrophosphate-dependent pump from the archaeon Pyrobaculum aerophilum: implications for the origins of pyrophosphate-energized pumps. FEBS Lett., 460: 505-512
Vatamaniuk, O.K., Mari, S., Lu, Y.-P., and Rea, P.A. (1999) AtPCS1, a phytochelatin synthase from Arabidopsis: isolation and in vitro reconstitution. Proc. Natl. Acad. Sci. USA., 96: 7110-7115
Rea, P.A. (1999) MRP subfamily ABC transporters from plants and yeast. J. Exp. Bot., 50: 895-913
Rea, P.A., Li, Z.-S., Lu, Y.-P., Drozdowicz, Y.M., and Martinoia, E. (1998) From vacuolar GS-X pumps to multispecific ABC transporters. Annu. Rev. Plant Physiol. Plant Mol. Biol., 49: 727-760
Lu, Y.-P., Li, Z.-S., Drozdowicz, Y.M., Hortensteiner, S., Martinoia, E., and Rea, P.A. (1998) AtMRP2, an Arabidopsis ATP-binding cassette transporter able to transport glutathione S-conjugates and chlorophyll catabolites: functional comparisons with AtMRP1. Plant Cell, 10: 1-18
Zhen, R.-G., Kim, E.J., and Rea, P.A. (1997) Acidic residues necessary for pyrophosphate-energized pumping and inhibition of the vacuolar proton-pyrophosphatase by N,N'-dicyclohexylcarbodiimide. J. Biol. Chem., 272: 22340-22348
Lu, Y.-P., Li, Z.-S., and Rea, P.A. (1997) AtMRP1 gene of A. thaliana encodes a glutathione S-conjugate pump: isolation and functional definition of a plant ATP binding cassette transporter gene. Proc. Natl. Acad. Sci. USA, 94: 8243-8248
Li, Z.-S., Lu, Y.-P., Thiele, D.J., and Rea, P.A. (1997) A new pathway for vacuolar cadmium sequestration in Saccharomyces cerevisiae: YCF1-mediated transport of bis(glutathionato)cadmium. Proc. Natl. Acad. Sci. USA, 94: 42-47
Zhen, R.-G., Kim, E.J., and Rea, P.A. (1997) The molecular and biochemical basis of pyrophosphate-energized proton translocation at the vacuolar membrane. Adv. Bot. Res., 25: 297-337.
Key Platform Publications:
Hirschi, K.D., Zhen, R.-G., Rea, P.A., and Fink, G.R. (1996) CAX1, an H+/Ca2+ antiporter from Arabidopsis. Proc. Natl. Acad. Sci. USA, 93: 8782-8786.
Li, Z.-S., Szczypka, M., Thiele, D.J., and Rea, P.A. (1996) The yeast cadmium factor protein (YCF1) is a vacuolar glutathione S-conjugate transporter. J. Biol. Chem., 271: 6509-6517.
Li, Z.-S., Zhen, R.-G., and Rea, P.A. (1995) 1-Chloro-2,4-dinitrobenzene-elicited increase in vacuolar MgATP-dependent glutathione S-conjugate transport. Plant Physiol., 109: 177-185
Li, Z.-S., Zhao, Y., and Rea, P.A. (1995) MgATP directly energizes glutathione S-conjugate transport by vacuolar membrane vesicles. Plant Physiol., 107: 1257-1268.
Kim, E.J., Zhen, R.-G., and Rea, P.A. (1995) Site-directed mutagenesis of vacuolar H+-pyrophosphatase: necessity of Cys634 for inhibition by maleimides but not catalysis. J. Biol. Chem., 270: 2630-2635.
Zhen, R.-G., Kim, E.J., and Rea, P.A. (1994) Localization of cytosolically oriented maleimide-reactive domain of vacuolar H+-pyrophosphatase. J. Biol. Chem.,, 269: 23342-23350.
Kim, Y., Kim, E.J., and Rea, P.A. (1994) Isolation and characterization of cDNAs encoding the vacuolar H+-pyrophosphatase of Beta vulgaris. Plant Physiol., 106: 373-382.
Kim, E.J., Zhen, R.-G., Rea, P.A. (1994) Heterologous expression of plant vacuolar pyrophosphatase in yeast demonstrates sufficiency of substrate-binding subunit for proton transport. Proc. Natl. Acad. Sci. USA, 91: 6128-6132.
Zhen, R.-G., Baykov, A.A., Bakuleva, N.P., Rea, P.A. (1994) Aminomethylenediphosphonate: a potent type-specific inhibitor of V-type H+-pyrophosphatases in plants and phototrophic bacteria. Plant Physiol., 104: 153-159.
Baykov, A.A., Kasho, V.N., Bakuleva, N.P., Rea, P.A. (1994) Oxygen exchange reactions catalyzed by vacuolar H+-translocating pyrophosphatase. Evidence for reversible formation of enzyme-bound pyrophosphate. FEBS Lett., 350: 323-327.
Baykov, A.A., Bakuleva, N.P., Rea, P.A. (1993) Steady state kinetics of substrate hydrolysis by vacuolar H+-pyrophosphatase: a simple three-state model. Eur. J. Biochem., 217: 755-762.
Baykov, A.A., Dubnova, E.B., Zhen, R.-G., Bakuleva, N.P., Evtushenko, O.A., Rea, P.A. (1993) Differential sensitivity of membrane-associated pyrophosphatases to diphosphonates and fluoride delineates two classes of enzyme. FEBS Lett., 317: 199-202.
Rea, P.A., Poole, R.J. (1993) Vacuolar H+-translocating inorganic pyrophosphatase. Annu. Rev. Plant Physiol. Plant Mol. Biol., 44: 157-180.
Davies, J.M., Poole, R.J., Rea, P.A., Sanders, D. (1992) Potassium transport into plant vacuoles is directly energized by a proton-pumping pyrophosphatase. Proc. Natl. Acad. Sci. USA, 89: 11701-11705.
Rea, P.A., Britten, C.J., Sarafian, V. (1992) Common identity of substrate-binding subunit of vacuolar H+-translocating inorganic pyrophosphatase of higher plants. Plant Physiol., 100: 723-732.
Britten, C.J., Zhen, R.-G., Kim, E.J., Rea, P.A. (1992) Reconstitution of transport function of vacuolar H+-translocating inorganic pyrophosphatase. J. Biol. Chem., 267: 21850-21855.
Rea, P.A., Britten, C.J., Jennings, I.R., Calvert, C.M., Skiera, L.A., Leigh, R.A., Sanders, D. (1992) Regulation of vacuolar H+-pyrophosphatase by free calcium: a reaction kinetic analysis. Plant Physiol., 100: 1706-1715.
Sarafian, V., Kim, Y., Poole, R.J., Rea, P.A. (1992) Molecular cloning and sequence of cDNA encoding the pyrophosphate-energized vacuolar membrane proton pump (H+-PPase) of Arabidopsis thaliana. Proc. Natl. Acad. Sci., USA, 89: 1775-1779.
Britten, C.J., Turner, J.C., Rea, P.A. (1989) Identification and purification of substrate binding subunit of higher plant H+-translocating inorganic pyrophosphatase. FEBS Lett., 256: 200-206.
Parry, R.V., Turner, J.C., Rea, P.A. (1989) High purity preparations of higher plant vacuolar H+-ATPase reveal additional subunits: revised subunit composition. J. Biol. Chem., 264: 20025-20032.
Rea, P.A., Sanders, D. (1987) Tonoplast energization: two H+ pumps, one membrane. Physiol. Plant., 71: 131-141.
Rea, P.A., Griffith, C.J., Sanders, D. (1987) Purification of N,N'-dicyclohexylcarbodiimide binding proteolipid of higher plant vacuolar H+-translocating ATPase. J. Biol. Chem., 262: 14745-14752.
Rea, P.A., Griffith, C.J., Manolson, M.F., Sanders, D. (1987) Irreversible inhibition of H+-ATPase of higher plant tonoplast by chaotropic anions: evidence for peripheral location of nucleotide-binding subunits. Biochim. Biophys. Acta, 904: 1-12.
Blumwald, E., Rea, P.A., Poole, R.J. (1987) Preparation of tonoplast vesicles: applications to H+-coupled secondary transport in plant vacuoles. Methods Enzymol., 148: 115-123.
Rea, P.A., Poole, R.J. (1986) Chromatographic resolution of H+-translocating inorganic pyrophosphatase from H+-translocating ATPase of higher plant tonoplast. Plant Physiol., 81: 126-129.
Manolson, M.F., Rea, P.A., Poole, R.J. (1985) Identification of BzATP- and DCCD-binding subunits of higher plant H+-translocating tonoplast ATPase. J. Biol. Chem., 260: 12273-12279.
Rea, P.A., Poole, R.J. (1985) Proton-translocating inorganic pyrophosphatase in red beet (Beta vulgaris L.) tonoplast vesicles. Plant Physiol., 77: 46-52.
American Association for the Advancement of Science; American Society for Biochemistry and Molecular Biology; American Society of Plant Biologists; Federation of American Societies for Experimental Biology; Magdalen Society, University of Oxford; New York Academy of Sciences